
Back مثيلة الحمض النووي الريبوزي المنقوص الأكسجين Arabic Metilacija DNK BS Metilació de l'ADN Catalan Methylace DNA Czech DNA-methylering Danish DNA-Methylierung German Metilación del ADN Spanish DNA metülatsioon Estonian متیلهشدن دیانای Persian Méthylation#Génétique French


DNA methylation is a biological process by which methyl groups are added to the DNA molecule. Methylation can change the activity of a DNA segment without changing the sequence. When located in a gene promoter, DNA methylation typically acts to repress gene transcription. In mammals, DNA methylation is essential for normal development and is associated with a number of key processes including genomic imprinting, X-chromosome inactivation, repression of transposable elements, aging, and carcinogenesis.
As of 2016, two nucleobases have been found on which natural, enzymatic DNA methylation takes place: adenine and cytosine. The modified bases are N6-methyladenine,[1] 5-methylcytosine[2] and N4-methylcytosine.[3]
| Unmodified base | 
|
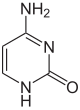
|
||||||
| Adenine, A | Cytosine, C | |||||||
| Modified forms | 
|

|

|
|||||
| N6-Methyladenine, 6mA | 5-Methylcytosine, 5mC | N4-Methylcytosine, 4mC | ||||||
Cytosine methylation is widespread in both eukaryotes and prokaryotes, even though the rate of cytosine DNA methylation can differ greatly between species: 14% of cytosines are methylated in Arabidopsis thaliana, 4% to 8% in Physarum,[4] 7.6% in Mus musculus, 2.3% in Escherichia coli, 0.03% in Drosophila; methylation is essentially undetectable in Dictyostelium;[5][6] and virtually absent (0.0002 to 0.0003%) from Caenorhabditis[7] or fungi such as Saccharomyces cerevisiae and S. pombe (but not N. crassa).[8][9]: 3699 Adenine methylation has been observed in bacterial and plant DNA, and recently also in mammalian DNA,[10][11] but has received considerably less attention.
Methylation of cytosine to form 5-methylcytosine occurs at the same 5 position on the pyrimidine ring where the DNA base thymine's methyl group is located; the same position distinguishes thymine from the analogous RNA base uracil, which has no methyl group. Spontaneous deamination of 5-methylcytosine converts it to thymine. This results in a T:G mismatch. Repair mechanisms then correct it back to the original C:G pair; alternatively, they may substitute A for G, turning the original C:G pair into a T:A pair, effectively changing a base and introducing a mutation. This misincorporated base will not be corrected during DNA replication as thymine is a DNA base. If the mismatch is not repaired and the cell enters the cell cycle the strand carrying the T will be complemented by an A in one of the daughter cells, such that the mutation becomes permanent. The near-universal use of thymine exclusively in DNA and uracil exclusively in RNA may have evolved as an error-control mechanism, to facilitate the removal of uracils generated by the spontaneous deamination of cytosine.[12] DNA methylation as well as a number of its contemporary DNA methyltransferases have been thought to evolve from early world primitive RNA methylation activity and is supported by several lines of evidence.[13]
In plants and other organisms, DNA methylation is found in three different sequence contexts: CG (or CpG), CHG or CHH (where H correspond to A, T or C). In mammals however, DNA methylation is almost exclusively found in CpG dinucleotides, with the cytosines on both strands being usually methylated. Non-CpG methylation can however be observed in embryonic stem cells,[14][15][16] and has also been indicated in neural development.[17] Furthermore, non-CpG methylation has also been observed in hematopoietic progenitor cells, and it occurred mainly in a CpApC sequence context.[18]
- ^ D. B. Dunn, J. D. Smith: "The occurrence of 6-methylaminopurine in deoxyribonucleic acids". In: Biochem J. 68(4), Apr 1958, S. 627–636. PMID 13522672. PMC 1200409.
- ^ B. F. Vanyushin, S. G. Tkacheva, A. N. Belozersky: "Rare bases in animal DNA". In: Nature. 225, 1970, S. 948–949. PMID 4391887.
- ^ Melanie Ehrlich, Miguel A. Gama-Sosa, Laura H. Carreira, Lars G. Ljungdahl, Kenneth C. Kuo, Charles W. Gehrke: "DNA methylation in thermophilic bacteria: N6-methylcytosine, 5-methylcytosine, and N6-methyladenine." In: Nucleic Acids Research. 13, 1985, S. 1399. PMID 4000939. PMC 341080.
- ^ Evans HH, Evans TE (December 1970). "Methylation of the deoxyribonucleic acid of Physarum polycephalum at various periods during the mitotic cycle". The Journal of Biological Chemistry. 245 (23): 6436–6441. doi:10.1016/S0021-9258(18)62627-4. PMID 5530731.

- ^ Smith SS, Rather DI (July 1991). "Lack of 5-methylcytosine in Dictyostelium discoideum DNA". The Biochemical Journal. 277 (1): 273–275. doi:10.1042/bj2770273. PMC 1151219. PMID 1713034.
- ^ Drewell RA, Cormier TC, Steenwyk JL, St Denis J, Tabima JF, Dresch JM, Larochelle DA (April 2023). "The Dictyostelium discoideum genome lacks significant DNA methylation and uncovers palindromic sequences as a source of false positives in bisulfite sequencing". NAR Genomics Bioinformatics. 5 (2): lqad035. doi:10.1093/nargab/lqad035. PMC 10111430. PMID 37081864.
- ^ Hu CW, Chen JL, Hsu YW, Yen CC, Chao MR (January 2015). "Trace analysis of methylated and hydroxymethylated cytosines in DNA by isotope-dilution LC-MS/MS: first evidence of DNA methylation in Caenorhabditis elegans". The Biochemical Journal. 465 (1): 39–47. doi:10.1042/bj20140844. PMID 25299492.
- ^ Bird A (December 2001). "Molecular biology. Methylation talk between histones and DNA". Science's Compass. Science. 294 (5549): 2113–2115. doi:10.1126/science.1066726. hdl:1842/464. PMID 11739943. S2CID 82947750.
As a result of this process, known as repeat-induced point mutation (RIP), the wild-type Neurospora genome contains a small fraction of methylated DNA, the majority of the DNA remaining nonmethylated.

- ^ Capuano F, Mülleder M, Kok R, Blom HJ, Ralser M (April 2014). "Cytosine DNA methylation is found in Drosophila melanogaster but absent in Saccharomyces cerevisiae, Schizosaccharomyces pombe, and other yeast species". Analytical Chemistry. 86 (8): 3697–3702. doi:10.1021/ac500447w. PMC 4006885. PMID 24640988.
- ^ Ratel D, Ravanat JL, Berger F, Wion D (March 2006). "N6-methyladenine: the other methylated base of DNA". BioEssays. 28 (3): 309–315. doi:10.1002/bies.20342. PMC 2754416. PMID 16479578.
- ^ Wu TP, Wang T, Seetin MG, Lai Y, Zhu S, Lin K, et al. (April 2016). "DNA methylation on N(6)-adenine in mammalian embryonic stem cells". Nature. 532 (7599): 329–333. Bibcode:2016Natur.532..329W. doi:10.1038/nature17640. PMC 4977844. PMID 27027282.
- ^ Angéla Békési and Beáta G Vértessy "Uracil in DNA: error or signal?"
- ^ Rana AK, Ankri S (2016). "Reviving the RNA World: An Insight into the Appearance of RNA Methyltransferases". Frontiers in Genetics. 7: 99. doi:10.3389/fgene.2016.00099. PMC 4893491. PMID 27375676.
- ^ Dodge JE, Ramsahoye BH, Wo ZG, Okano M, Li E (May 2002). "De novo methylation of MMLV provirus in embryonic stem cells: CpG versus non-CpG methylation". Gene. 289 (1–2): 41–48. doi:10.1016/S0378-1119(02)00469-9. PMID 12036582.
- ^ Haines TR, Rodenhiser DI, Ainsworth PJ (December 2001). "Allele-specific non-CpG methylation of the Nf1 gene during early mouse development". Developmental Biology. 240 (2): 585–598. doi:10.1006/dbio.2001.0504. PMID 11784085.
- ^ Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, et al. (November 2009). "Human DNA methylomes at base resolution show widespread epigenomic differences". Nature. 462 (7271): 315–322. Bibcode:2009Natur.462..315L. doi:10.1038/nature08514. PMC 2857523. PMID 19829295.
- ^ Lister R, Mukamel EA, Nery JR, Urich M, Puddifoot CA, Johnson ND, et al. (August 2013). "Global epigenomic reconfiguration during mammalian brain development". Science. 341 (6146): 1237905. doi:10.1126/science.1237905. PMC 3785061. PMID 23828890.
- ^ Kulis M, Merkel A, Heath S, Queirós AC, Schuyler RP, Castellano G, et al. (July 2015). "Whole-genome fingerprint of the DNA methylome during human B cell differentiation". Nature Genetics. 47 (7): 746–756. doi:10.1038/ng.3291. PMC 5444519. PMID 26053498.
© MMXXIII Rich X Search. We shall prevail. All rights reserved. Rich X Search