
Back إنزيم مدمج Arabic ইনটিগ্রেজ Bengali/Bangla Integrasa Catalan Integráza Czech Integrase German Integrasa Spanish Integrasa Basque اینتگراز Persian Intégrase French Integrasi Italian
| Integrase Zinc binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 solution structure of the n-terminal zn binding domain of hiv-1 integrase (e form), nmr, 38 structures | |||||||||
| Identifiers | |||||||||
| Symbol | Integrase_Zn | ||||||||
| Pfam | PF02022 | ||||||||
| InterPro | IPR003308 | ||||||||
| SCOP2 | 1wjb / SCOPe / SUPFAM | ||||||||
| |||||||||
| Integrase core domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
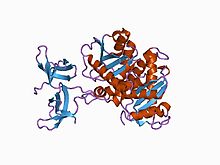 Crystal structure of the RSV two-domain integrase. | |||||||||
| Identifiers | |||||||||
| Symbol | rve | ||||||||
| Pfam | PF00665 | ||||||||
| Pfam clan | CL0219 | ||||||||
| InterPro | IPR001584 | ||||||||
| SCOP2 | 2itg / SCOPe / SUPFAM | ||||||||
| |||||||||
| Integrase DNA binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Crystal structure of the RSV two-domain integrase. | |||||||||
| Identifiers | |||||||||
| Symbol | IN_DBD_C | ||||||||
| Pfam | PF00552 | ||||||||
| InterPro | IPR001037 | ||||||||
| SCOP2 | 1ihw / SCOPe / SUPFAM | ||||||||
| |||||||||
Retroviral integrase (IN) is an enzyme produced by a retrovirus (such as HIV) that integrates (forms covalent links between) its genetic information into that of the host cell it infects.[1] Retroviral INs are not to be confused with phage integrases (recombinases) used in biotechnology, such as λ phage integrase, as discussed in site-specific recombination.
The macromolecular complex of an IN macromolecule bound to the ends of the viral DNA ends has been referred to as the intasome; IN is a key component in this and the retroviral pre-integration complex.[2]
- ^ Beck BJ, Freudenreich O, Worth JL (2010). "Patients with Human Immunodeficiency Virus Infection and Acquired Immunodeficiency Syndrome". Massachusetts General Hospital Handbook of General Hospital Psychiatry. Elsevier. pp. 353–370. doi:10.1016/b978-1-4377-1927-7.00026-1. ISBN 9781437719277.
- ^ Masuda T (2011). "Non-Enzymatic Functions of Retroviral Integrase: The Next Target for Novel Anti-HIV Drug Development". Frontiers in Microbiology. 2: 210. doi:10.3389/fmicb.2011.00210. PMC 3192317. PMID 22016749.
© MMXXIII Rich X Search. We shall prevail. All rights reserved. Rich X Search